The progenitor organism from which all current life forms ultimately descended did not materialize on Earth abruptly approximately 4.2 billion years ago.
A portion of its genetic material originated from an even more ancient and enigmatic source…
“While the most ancient organism discernible through evolutionary methodologies is the last universal common ancestor,” elaborates Oberlin College biologist Aaron Goldman, “certain genes within its blueprint were considerably older.”
These exceptionally ancient protein-coding genes warrant deeper investigation if our objective is to illuminate the fundamental origins of life on our planet, argue Goldman, alongside MIT’s Greg Fournier and University of Wisconsin-Madison’s Betül Kaçar, in a recent publication.
Though the practice of inferring the earliest epochs of evolutionary history using ancient gene families is not novel, this scientific cohort aims to emphasize its significance now that pivotal advancements in ancestral sequence reconstruction have enabled unprecedented scrutiny of the genome of the last universal common ancestor (LUCA).
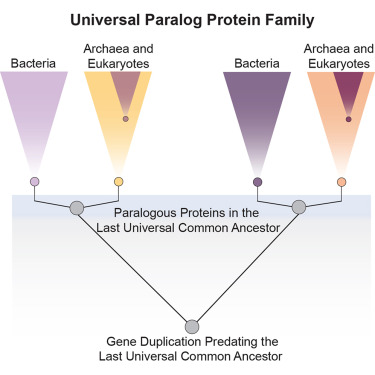
Genetics, akin to historical narratives, is predominantly shaped by enduring lineages. Organisms that fail to propagate descendants leave minimal trace of their existence.
The geological strata do not extend back sufficiently far to encompass LUCA’s probable epoch, rendering our genetic makeup one of the few tangible remnants of that era.
Gene families that underwent ancient duplication, also referred to as ‘universal paralogs’, represent infrequent instances of gene redundancy present across every extant life form. This implies their duplication occurred prior to the divergence of these lineages.
If LUCA serves as the central nexus of our genetic lineage, then the primordial unicellular organisms that likely possessed these duplicated genes constitute the very foundations of life – the deeply embedded precursors that facilitated the emergence of animals, plants, fungi, and bacteria.
“By following the trajectory of universal paralogs,” states Kaçar, “we can bridge the initial stages of terrestrial life with contemporary scientific instrumentation. These offer an opportunity to transform the most profound enigmas in evolution and biology into empirically verifiable discoveries.”
Scientists can only formulate speculative hypotheses regarding the conditions during LUCA’s existence, let alone the events preceding it. It is plausible that LUCA did not inhabit Earth in isolation but rather existed within an “established ecological framework” characterized by moderate productivity.
The degree of simplicity or complexity inherent in these organisms or their ecosystems remains a subject of ongoing discussion.
Currently, only a limited number of universal paralog protein families are recognized, the authors Goldman and his collaborators report, but this scarcity “does not necessarily indicate a deficit of paralogs within LUCA’s actual proteome.”
Over extended periods, a multitude of paralogs within LUCA’s proteome were likely eliminated from the evolutionary record due to various factors, including evolutionary shifts, genetic divergence, or horizontal gene transfer (a prevalent mechanism for gene exchange among bacterial populations).
Such events could obscure the ancient origins of certain protein-specific genes that persist in modern biology.
“Consequently, it is probable that a majority of protein families present in LUCA might not be discernible through phylogenetic analyses,” the researchers concur.
While this presents a constraint, it correspondingly enhances the significance of any discoverable elements.
“The historical record of these universal paralogs represents the sole repository of information concerning these nascent cellular lineages, necessitating that we meticulously extract every available insight,” emphasizes Fournier.
For instance, some universal paralogs are integral to the genetic translation machinery, which the perspective authors posit “is very likely the most ancient molecular system preserved in extant life today.”
Additional universal paralogs are implicated in the synthesis of enzymes or in proteins crucial for maintaining the integrity of biological membranes.
Recent investigations, for example, have illuminated the existence of pre-LUCA progenitors for enzymes known as aminoacyl tRNA synthetases, as documented in a body of research.
Describing these enzymes as vital to life would be an understatement. Their function involves the precise attachment of the correct amino acid to its cognate transfer RNA, which subsequently facilitates the sequential arrangement of these amino acids to construct a protein.
The presence of their ancestors preceding LUCA suggests that early life forms possessed the capacity to incorporate amino acids into genetically encoded proteins even prior to the evolution of their more contemporary counterparts.
“In summation, these findings corroborate the notion that the evolution of a modern genetic code before LUCA constituted a multifaceted developmental trajectory involving numerous mechanisms, including co-evolution with amino acid biosynthesis pathways,” the authors conclude.
The findings of this study have been published in the journal Cell Genomics.