Researchers at the University of California, Santa Barbara, led by Professor Soojin Yi, embarked on a mission to investigate the evolutionary trajectory of genes within distinct brain cell populations, comparing them to those found in chimpanzees. Their inquiry yielded a significant discovery: while our genetic makeup encodes for a nearly identical set of proteins as other great apes, a substantial number of our genes exhibit considerably higher productivity than their counterparts in other primates.
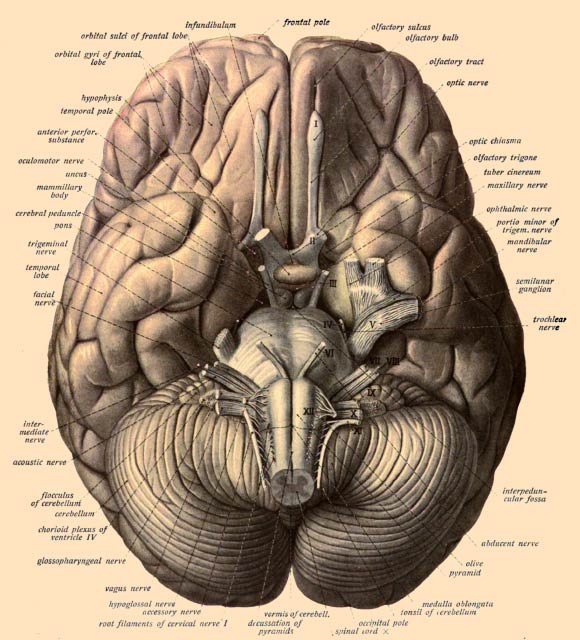
Differential gene expression in the human brain compared to nonhuman primate brains is a key molecular feature of human evolution. Joshy et al. show that differential gene expression of human brains encompasses highly divergent sets of cell-type-specific changes. Despite such cellular diversity, human brain cells have experienced a general increase of gene expression rather than a decrease of gene expression. The authors reveal specific functional programs that have experienced differential expression in different cell types and genomic and epigenomic features that correlate with such changes.
As the scientific community began to grasp the fundamental role of the genome as life’s foundational blueprint, the hypothesis emerged that our own genome might hold the key to explaining uniquely human characteristics.
However, an exhaustive comparative analysis with chimpanzees conducted in 2005 revealed a striking genetic similarity, with humans sharing approximately 99% of their genes—a figure that has since been subject to revision by scientific consensus.
This finding corroborated earlier investigations that, based on limited gene sampling, had suggested minimal genetic divergence between the human and chimpanzee genomes.
Contemporary biological understanding now posits that differences in gene expression may be the underlying factor for these observed distinctions. To illustrate, consider the metamorphosis of a monarch butterfly: the adult insect possesses the identical genome as its larval caterpillar stage. The profound transformations between these life phases are entirely attributed to variations in gene expression. The selective activation or silencing of specific genes, or their capacity to synthesize varying quantities of messenger RNA (mRNA), can lead to dramatic alterations in an organism’s phenotypic traits.
Illuminating the Landscape
Prior research had identified disparities in gene expression patterns between humans and chimpanzees, indicating a general tendency towards elevated gene expression in human cells. Nevertheless, the overall picture remained somewhat indistinct.
The brain, a remarkably complex organ, is comprised of a multitude of distinct cell types.
Historically, neuroscience has broadly categorized brain cells into two primary classifications: neurons and glial cells.
Neurons are instrumental in transmitting electrochemical signals, functioning analogously to the electrical wiring within a structure.
Glial cells, on the other hand, undertake a diverse array of supporting roles, including the insulation of neural pathways, structural reinforcement, and the removal of cellular waste products.
Until a recent technological advancement, scientific inquiry was limited to the analysis of bulk tissue samples, which contained a heterogeneous mix of various cell types. However, within the last decade, the capability to single-cell analysis of cellular nuclei has become attainable.
This breakthrough empowers researchers to differentiate between distinct cell types and, with enhanced precision, even identify specific subtypes.
Professor Yi and her collaborators leveraged datasets meticulously compiled using a specialized device equipped with a microfluidic channel, enabling the isolation of individual nuclei into separate compartments within an array.
Subsequently, these cells were meticulously categorized by type prior to undergoing rigorous statistical examination.
Gene expression levels were quantified by measuring the quantity of mRNA produced by a particular gene within humans, chimpanzees, and macaques.
A gene identified as upregulated demonstrates a greater mRNA output in a given species relative to others, whereas a downregulated gene exhibits a reduced mRNA production.
By conducting comparative analyses between chimpanzees and humans, utilizing macaques as an evolutionary reference point, the researchers were equipped to discern whether observed differences between the two ape species stemmed from evolutionary changes within the chimpanzee lineage, the human lineage, or a combination of both.
The study’s authors cataloged variations in the expression of approximately 5-10% of the 25,000 genes analyzed.
Broadly speaking, human cells exhibited a greater prevalence of upregulated genes when contrasted with chimpanzee cells.
This represents a considerably larger proportion than was detected in previous studies that did not stratify the analysis by cell type. Furthermore, this percentage escalated to 12-15% when the authors incorporated the analysis of cell subtypes.
“We are now able to observe that individual cell types possess their own unique evolutionary trajectories, leading to remarkable specialization,” Professor Yi remarked.
Beyond Neuronal Dominance
The intricate complexity of human neural circuitry is unparalleled across the animal kingdom. Nevertheless, the research team hypothesizes that our distinctive cognitive abilities are not solely attributable to this factor.
In the human brain, glial cells constitute over half of the total cellular population, a significantly higher proportion than observed even in chimpanzees.
Among the various types of glial cells, oligodendrocytes displayed the most pronounced discrepancies in gene expression. These cells are responsible for generating the myelin sheath that insulates neurons, thereby facilitating the significantly faster and more efficient transmission of electrical signals.
In a collaborative research endeavor published in the preceding year, the scientists noted a higher ratio of precursor oligodendrocytes to mature oligodendrocytes in humans compared to chimpanzees.
This observation leads them to suspect a potential correlation with the remarkable neural plasticity and prolonged developmental period characteristic of human brains.
“The enhanced complexity of our neural network likely did not evolve in isolation,” Professor Yi stated.
“Its emergence would have been contingent upon the co-evolution of all these ancillary cell types, which in turn facilitated the expansion of neuronal diversity, the sheer number of neurons, and the intricate architecture of the neural networks.”
The findings of this research have been formally published in the esteemed journal Proceedings of the National Academy of Sciences.
_____
Dennis Joshy et al. 2024. Accelerated cell-type-specific regulatory evolution of the human brain. PNAS 121 (52): e2411918121; doi: 10.1073/pnas.2411918121
This article has been adapted from an original dissemination by the University of California, Santa Barbara.





